Fatty acids, the compounds that give a diet rich in leafy greens and fish its antioxidant and anti-inflammatory benefits, are now also heralded for their versatility as raw materials in bioenergy production.
Great Lakes Bioenergy Research Center scientists are enamored with one particular kind of long chain fatty acid, called furan fatty acid, because it could substitute for petroleum-based products including fuel, engine lubricant, medicines and food additives. Now, a team of GLBRC collaborators at the University of Wisconsin–Madison have described a pathway for furan fatty acid production in bacteria and other cells.
Similar to other fatty acids, furan fatty acids, or FuFAs, are found in the membrane that forms a cell’s border. These fatty acids act like an oily filter to protect the interior of the cell against changes in the external environment. They can also act as chemical messengers that tell the cell when a toxin or stress condition is present.
FuFAs are a special class of fatty acids with broad appeal to biofuel scientists because an oxygen atom is attached in the middle of the hydrocarbon chain.

“The chemistry of FuFAs helps the bacterial cell fight off damage in the membrane. The oxygen atom in FuFAs is also what makes it interesting to the biofuel and other industries,” says Rachelle Lemke, a GLBRC senior research specialist who led a study of FuFA production published recently in the Journal of Biological Chemistry. “Think of oxygen as the lighter that helps ignite all the other molecules in a fuel to burn. The chain length of FuFAs also make it a potential lubricant for engines and other devices.”
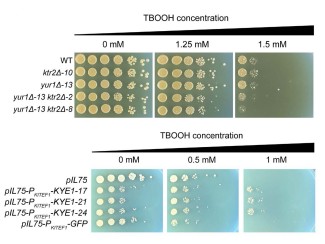
Lemke previously identified FuFAs while exploring why a bacterium, Rhodobacter spheroides, dies when exposed to a hyperreactive oxygen molecule called singlet oxygen. As a graduate student studying with Tim Donohue, UW–Madison bacteriology professor and GLBRC director, Lemke originally found that singlet oxygen kills R. spheroides unless FuFAs are present. Still, they lacked information on how these important class of fatty acids were made.
In their new study, Lemke and a team of GLBRC researchers identified the steps this and other bacteria use to produce FuFAs. The team’s findings reveal previously unidentified intermediate molecules that ultimately get converted into FuFA. They also found enzymes that add oxygen to the FuFA chain while decorating the ring with methyl groups that serve as the chemical makeup for the genesis of fuels.

The team also used genome sequences and genetics to identify enzymes that perform chemical reactions that have never been observed until now. With these genetic GPS signals, the team was able to identify proteins that catalyze the same reactions in one other bacterium and predict the presence of this newly described pathway for FuFA production in both microbes and plants.
The next steps for GLBRC scientists are to understand the molecular basis of these newly discovered enzymes and to train cells to make more FuFAs so researchers can test the fatty acids in a variety of applications.
“With more tests, it’s possible we will understand how the new classes of enzymes work, and design cells to crank up FuFA production in bacteria and other cells,” Lemke says.
What began more than a decade ago with a simple question: “why do some bacteria die when there’s oxygen around?” has added another layer to the production line of fuels and bioproducts from plants, now patented and available for licensing through the Wisconsin Alumni Research Foundation.
“This discovery is another great example about how interdisciplinary GLBRC teams can mine genomes to reveal previously unknown biochemical pathways,” says Donohue. “In addition, we have received interest in evaluating furan fatty acids as lubricants, as compounds with antimicrobial activity and as food additives from groups around the globe.”
GLBRC is one of four U.S. Department of Energy Bioenergy Research Centers created to provide scientific breakthroughs for a new generation of sustainable, cost-effective biofuels and bioproducts. For more information on GLBRC, visit www.glbrc.org or visit us on twitter.
Publication
A bacterial biosynthetic pathway for methylated furan fatty acids
Rachelle A. S. Lemke, Stephanie M. Olson, Kaitlin Morse, Steven D. Karlen, Alan Higbee, Emily T. Beebe, John Ralph, Joshua J. Coon, Brian G. Fox and Timothy J. Donohue
DOI: 10.1074/jbc.RA120.013697
Lemke, R.A.S., et al. “A bacterial biosynthetic pathway for methylated furan fatty acids.” Journal of Biological Chemistry 295, 9786-9801 (2020). [DOI: 10.1074/jbc.RA120.013697]