Study reveals link between specificity and efficiency in molecular machines that could be engineered to improve biofuel production
The Science
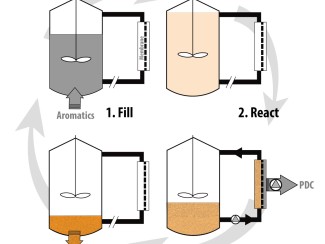
Membrane transporters are corkscrew-shaped proteins that living cells use to get rid of toxins and other harmful substances. Also called efflux pumps, these molecular machines serve as tiny gatekeepers, allowing only certain molecules to pass through the cell membrane. Efflux pumps are a problem in medicine because they enable microbes to expel drugs that are supposed to kill them, which leads to antibiotic resistance. But efflux pumps present an opportunity to improve biomanufacturing if they can help cells get rid of toxins that slow fermentation. Whether the goal is modifying transporters to prevent antibiotics from escaping the cell or to remove fermentation inhibitors, scientists need to understand how these proteins form and how they interact with various molecules.
Some transporters can expel more than one type of substance, raising questions about how they are able to recognize molecules with different structures while still being selective and whether there is a connection between a pump’s breadth and the energy required to operate it. To study that relationship, researchers grew bacteria with thousands of variations of a transporter from a microbe responsible for thousands of deaths each year. They then analyzed how each mutation affected the pump's selectivity and energy efficiency. The results showed a direct relationship between energy use and breadth: more efficient pumps can generally transport a wider array of substances, while inefficient variants tend to have a narrower profile.
The Impact
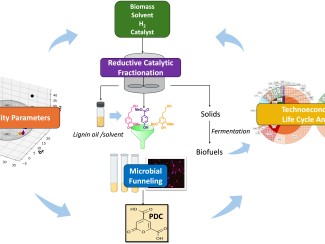
Bacteria and other microbes can convert plant fibers into sustainable fuels and chemicals used to make plastics, medicines, and other products. But chemicals used in processing or in the plants themselves are an obstacle because they can kill the cells or slow fermentation. Researchers are looking for ways to modify natural efflux pumps to selectively remove these toxins, but testing the vast number of possible variations is impractical using traditional lab techniques. Data generated for this project are being used to train artificial intelligence models to predict which mutations are most likely to be effective.
Summary
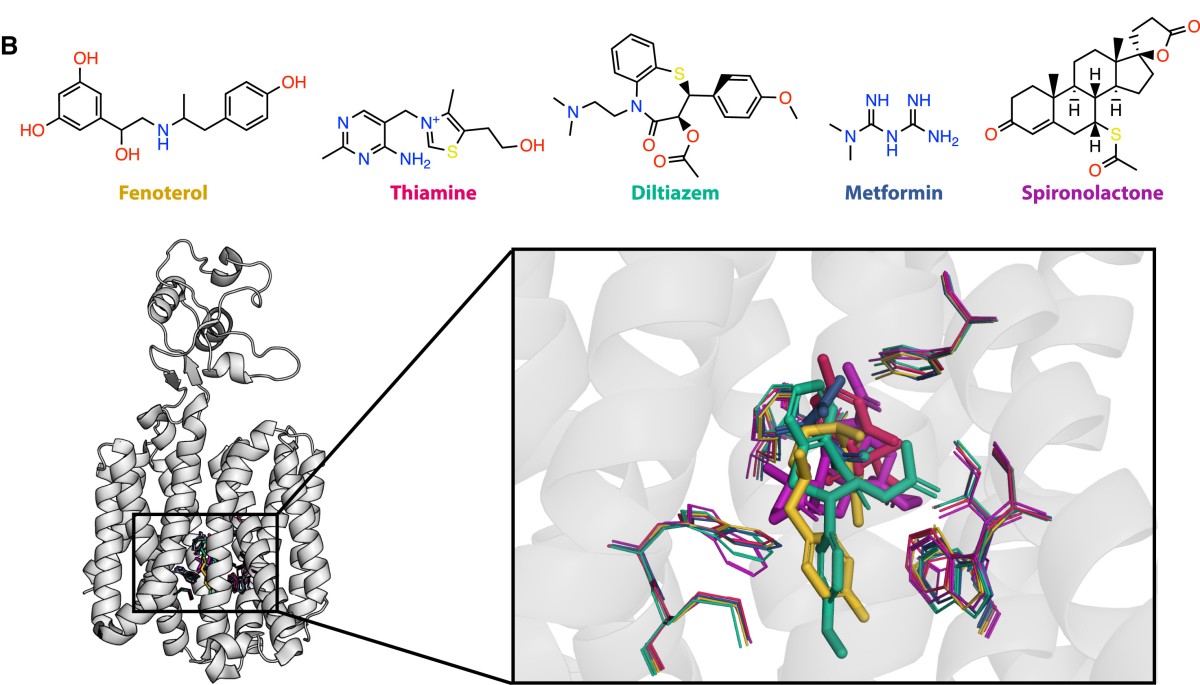
Scientists with the Great Lakes Bioenergy Research Center used deep mutational scanning across eight diverse chemicals (substrates) and two energy conditions to analyze the NorA efflux pump, a proton-coupled transporter of Staphylococcus aureus that confers resistance to a range of antibiotics and is frequently overexpressed in multidrug resistance infections. They developed multiparametric screens to deconvolute functional effects into distinct elements of fitness: drug resistance, energy efficiency, and stability.
Data revealed that substrate specificity is not confined to the binding site, but arises from a distributed network of residues throughout the protein. Measuring pH-dependent transport efficiency as a proxy for energy coupling revealed a fundamental coupling between energy utilization and substrate breadth: Efficient variants maintain broad specificity, while inefficient variants are narrower. They developed a thermodynamic framework that links energy efficiency to substrate promiscuity. Together the findings establish fundamental principles of transporter polyspecificity and provide a blueprint for understanding, predicting, and engineering substrate selectivity.