Some chemicals used to speed up the breakdown of plants for production of biofuels like ethanol are poison to the yeasts that turn the plant sugars into fuel.
Researchers from the University of Wisconsin–Madison and several Department of Energy laboratories have identified two changes to a single gene that can make the yeast tolerate the pretreatment chemicals. They published their findings recently in the journal Genetics.
Even in a cutting-edge factory, turning plant material like grasses or leftover cornstalks into biofuels often mimics the way nature returns plant nutrients to the soil, air and water. One way or another, the plant cells are broken down physically, chemically and with microbes, almost like it would decompose naturally.
“But the process of decomposing plant material is really slow. It takes years for a fallen tree to completely decompose,” says Trey Sato, a senior scientist at the UW–Madison-based Great Lakes Bioenergy Research Centerand a lead researcher on the yeast study. “That length of time isn’t compatible with industrial situations, where the goal is to make as much product as fast as possible in order to get it to market for sale.”
So biofuels manufacturers speed up the process in part by pretreating the raw plant biomass. Pretreatment may include applying ammonia gas, acids, heat and pressure, salts called ionic liquids, or some combination of those and other schemes.
After pretreatment, the cellulose that makes up the plant cell walls and fibers is broken down with enzymes to release sugar. The sugar is fermented into fuel by microbes — often carefully bred and engineered versions of the yeast Saccharomyces cerevisiae, also used to ferment wine and beer and leaven bread.
“Those ionic liquids are useful for pretreating and getting the process started,” Sato says. “The problem is that even after you go to the trouble to remove and recover as much of the ionic liquids as you can from your biomass before you do the fermenting, the amount you can’t get out is enough to be toxic to a lot of microbes.”
This toxicity is enough to make the yeast as much as 70 percent less efficient at turning sugar into biofuel, a crippling loss for an industrial process.
Sato and collaborators at UW–Madison, the Joint BioEnergy Institute and Lawrence Livermore, Lawrence Berkeley and Sandia national laboratories went looking for a way to overcome this barrier.
“Nature is probably the best engineer there is. It’s had millions of years of evolution to develop and optimize biological systems,” Sato says. “So one solution might be to go out and find a replacement for yeast, a different microbe that might be able to ferment cellulosic sugars and also stomach the ionic liquids.”
But many biofuels companies and engineers have invested a great deal of time and resources in tweaking their own yeast and building biofuel reactors around their preferred strains. A different fermenting organism might put them back at square one.
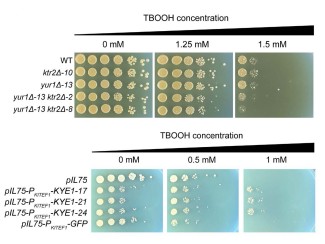
So, the researchers looked to a variety of S. cerevisiae strains that were isolated from different ecological niches. Of the 136 yeast isolates they surveyed, they found one strain with outstanding tolerance to ionic liquids. They screened DNA sequences from this strain and identified a pair of genes key to surviving the otherwise toxic pretreatment chemicals. One of the genes, called SGE1, makes a protein that settles in the yeast cell membrane and works as a pump to remove toxins.
“If you have more of these pumps at the cell surface, you can get more of the ionic liquid molecules out of your cell,” Sato says.
The researchers found a strain of yeast with outstanding tolerance to ionic liquids, and identified a gene that makes a protein that works as a pump to remove toxins.
A change of just two individual nucleotides among more than 12 million that make up the yeast genome are enough to increase the production of those cellular pumps and protect yeast from ionic liquids. The researchers used the gene-editing tool CRISPR to alter a strain of an ionic liquid-susceptible yeast, introducing the two single-nucleotide changes and successfully producing a yeast that can survive — and ferment — alongside amounts of ionic liquid that are normally toxic.
“Now anyone using this yeast can look at a specific gene in their own strain and tell whether it’s compatible and useful with an ionic liquid process or not,” says Sato. “It’s a simple engineering procedure, which doesn’t take long and isn’t expensive. And it can be fixed with CRISPR in a matter of a week or two.”
Sato says the next step is to try out the modified yeast outside the lab, incorporating the real-world plant material used as biofuel feedstock.
Collaborators on the study included Michael Thelen of the Joint BioEnergy Institute and UW–Madison genetics professors Audrey Gasch and Chris Hittinger and biochemistry and bacteriology professor Robert Landick.
THIS RESEARCH WAS SUPPORTED BY GRANTS FROM THE DEPARTMENT OF ENERGY (DE-SC0018409, DE-FC02-07ER64494 AND DE-AC02-05CH11231).
Abstract
Douglas A. Higgins, Megan K. Young, Mary Tremaine, Maria Sardi, Jenna M. Fletcher, Margaret Agnew, Lisa Liu, Quinn Dickinson, David Peris, Russell L. Wrobel, Chris Todd Hittinger, Audrey P. Gasch, Steven W. Singer, Blake A. Simmons, Robert Landick, Michael P. Thelen and Trey K. Sato
GENETICS Early online July 25, 2018; https://doi.org/10.1534/genetics.118.301161
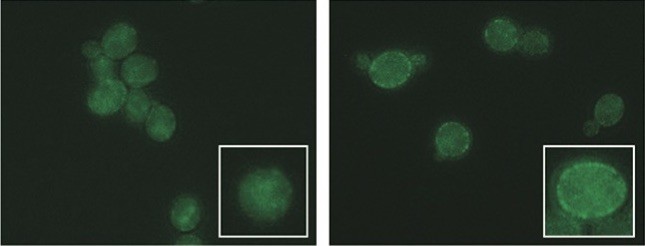
Imidazolium ionic liquids (IILs) have a range of biotechnological applications, including as pretreatment solvents that extract cellulose from plant biomass for microbial fermentation into sustainable bioenergy. However, residual levels of IILs, such as 1-ethyl-3-methyl imidazolium chloride ([C2C1im]Cl), are toxic to biofuel-producing microbes, including the yeast Saccharomyces cerevisiae. S. cerevisiae strains isolated from diverse ecological niches differ in genomic sequence and in phenotypes potentially beneficial for industrial applications, including tolerance to inhibitory compounds present in hydrolyzed plant feedstocks. We evaluated over one-hundred genome-sequenced S. cerevisiae strains for tolerance to [C2C1im]Cl and identified one strain with exceptional tolerance. By screening a library of genomic DNA fragments from the [C2C1im]Cl-tolerant strain for improved IIL tolerance, we identified SGE1, which encodes a plasma membrane multidrug efflux pump, and a previously uncharacterized gene that we named ILT1, which encodes a predicted membrane protein. Analyses of SGE1sequences from our panel of S. cerevisiae strains together with growth phenotypes implicated two single nucleotide polymorphisms (SNPs) that associated with IIL tolerance and sensitivity. We confirmed these phenotypic effects by transferring the SGE1 SNPs into a [C2C1im]Cl-sensitive yeast strain using CRISPR/Cas9 genome editing. Further studies indicated that these SNPs affect Sge1 protein stability and cell surface localization, impacting the amount of toxic IILs that cells can pump out of the cytoplasm. Our results highlight the general potential for discovering useful biotechnological functions from untapped natural sequence variation and provide functional insight into emergent SGE1 alleles with reduced capacities to protect against IIL toxicity.