Better understanding of the family responsible for nearly all bread, fermented drinks, and biofuels creates a path to finding new strains with potentially useful traits.
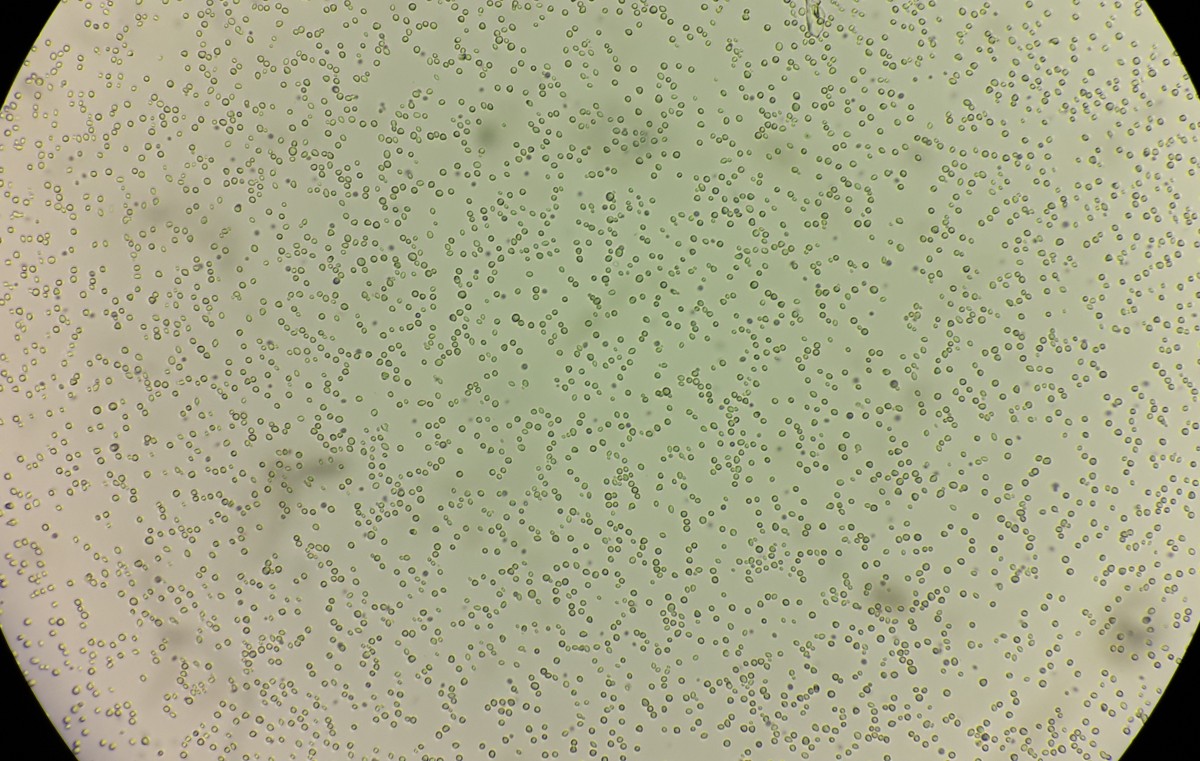
Commonly known as brewer’s yeast, Saccharomyces cerevisiae tends to get credit for bread, beer, wine, ethanol and just about any other product of fermentation.
In fact, for many products, the bulk of the work is done by interspecies hybrids, strains crossbred with other yeast species. Yet most scientific study of the genus has focused on domesticated strains of S. cerevisiae, limiting understanding of how these fungi evolved over the past 20 million years and how other varieties could be used to solve challenges like converting plants into sustainable fuels and other products.
Now, an international team led by scientists with the Great Lakes Bioenergy Research Center at UW–Madison have analyzed hundreds of yeasts, including six little-studied wild species, in the most comprehensive study to date of the Saccharomyces genus.
The results, published this spring in the journal Nature Communications, create a framework for understanding biodiversity in microbial organisms and their potential uses.
Lead author David Peris said the results show how microscopic organisms found in forests and parks around the world can be used to make better beer, wine, bread, and plant-based replacements for fossil fuels and other petrochemicals.
“We have discovered new strains with interesting industrial traits,” Peris said.
For example, S. mikatae, a species never used in beer fermentation, actually grows in maltotriose, a sugar found in the raw material used to make beer.

Peris and his collaborators examined more than 1,800 strains of Saccharomyces collected from soils and tree bark on every continent but Antarctica and Africa, cataloging observable traits and genetic variations to better understand how these yeasts evolved over millions of years.
The researchers generated and analyzed complete genome sequences – the biological blueprints that guide how organisms develop and function – of 163 strains and cataloged the observable traits of 128 strains from different lineages.
They also quantified diversity and differences within and between species and populations, as well as natural reticulation events, such as interbreeding, the influences of ecology, and genetic differences that don’t match the expected pattern.
This type of work is key to the Biden administration’s ambitious biotech goals, which include replacing up to 90% of plastics with biobased processes in the next two decades. The plan calls for sequencing the genomes of a million microbial species to better understand the function of these newly discovered genes.
Peris, now a researcher with the Spanish National Research Council’s Institute of Agrochemistry and Food Technology and the University of Oslo in Norway, began the work in 2012 as a postdoctoral researcher in UW–Madison genetics professor Chris Todd Hittinger’s lab.
Hittinger, whose team has discovered thousands of new wild strains, said the study of yeasts has historically been biased toward human culture. But over the past decade or so, he and other researchers began collecting wild yeasts, which they isolated in the lab to form a sort of “museum of microbes.”
The goal of this study was to gain a better understanding of those wild yeasts, with an eye toward harnessing their traits to benefit humans.
“We have a large collection of strains from other species in the same genus, but people were not exploring them,” Peris said. “We didn’t know their genome or traits. For example, could they survive in [a particular] environment? And could those traits be of interest to industry?”
By cross-breeding species with different attributes, scientists can bioengineer yeasts that are better at certain jobs. For example, strains of yeast with faster metabolisms could reduce the time required for fermentation, cutting production costs. Others might be better able to tolerate the higher temperatures or toxins associated with production of biofuels like ethanol or isobutanol.

In order to do that, scientists need to understand the genetic code that makes those yeasts tick, a task that would have been nearly impossible a couple of decades ago.
Hittinger compares the process to the parable of the elephant and the blind men.
“Different parts of the genome have different histories,” Hittinger said. “So before next-generation sequencing technologies lowered the cost of genome sequencing, researchers would sometimes come to different conclusions depending on which part of the genome they examined.”
But technology developed in the late 2000s made it possible to sequence a genome in just a few days for less than $100, a feat that in 1996 took an international team of scientists several years and cost millions of dollars.
“This is really a case where it took both innovation on the natural history side -- pulling yeasts out of nature -- and the new technology of next-generation sequencing,” Hittinger said.
Peris said his work on the genus Saccharomyces can now serve as a model for understanding and delineating lineages, which scientists can use as they explore other groups of yeasts.
He and Hittinger are continuing their collaboration on a project called Sacchar2Omics that aims to sequence additional strains of Saccharomyces in hopes of finding other traits that could aid in the production of sustainable foods, biofuels, and other plant-based products.
Meanwhile Hittinger is leading a five-year project to chart the evolution of more than a thousand yeast species over the past 400 million years, the first step toward harnessing their ability to metabolize carbon sources.